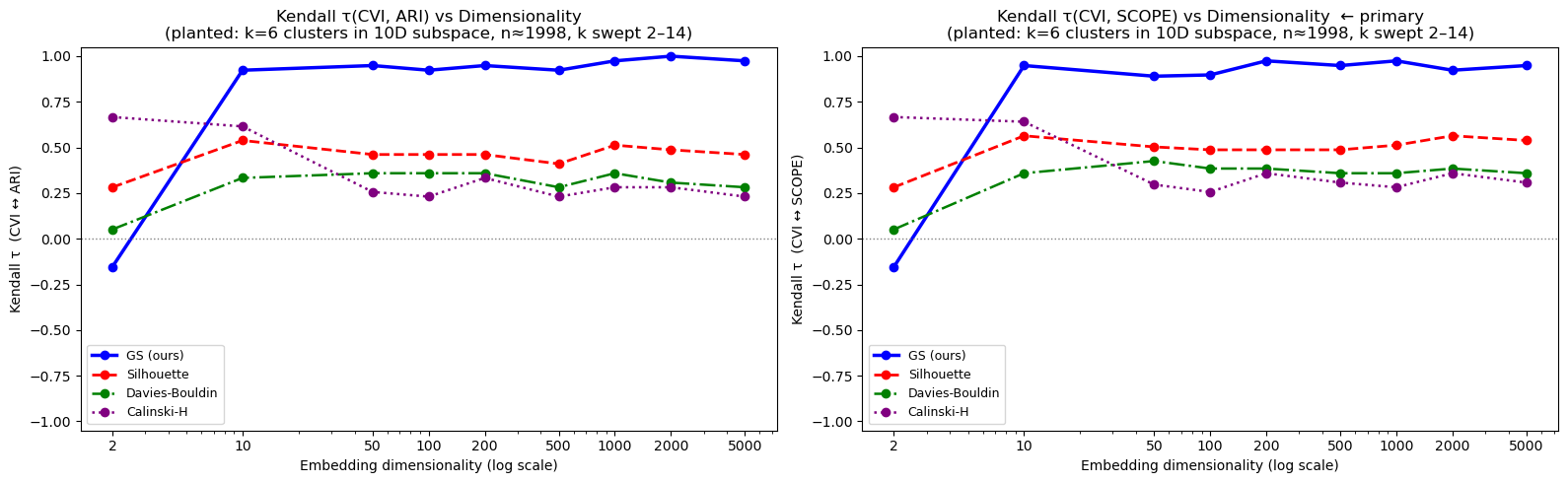
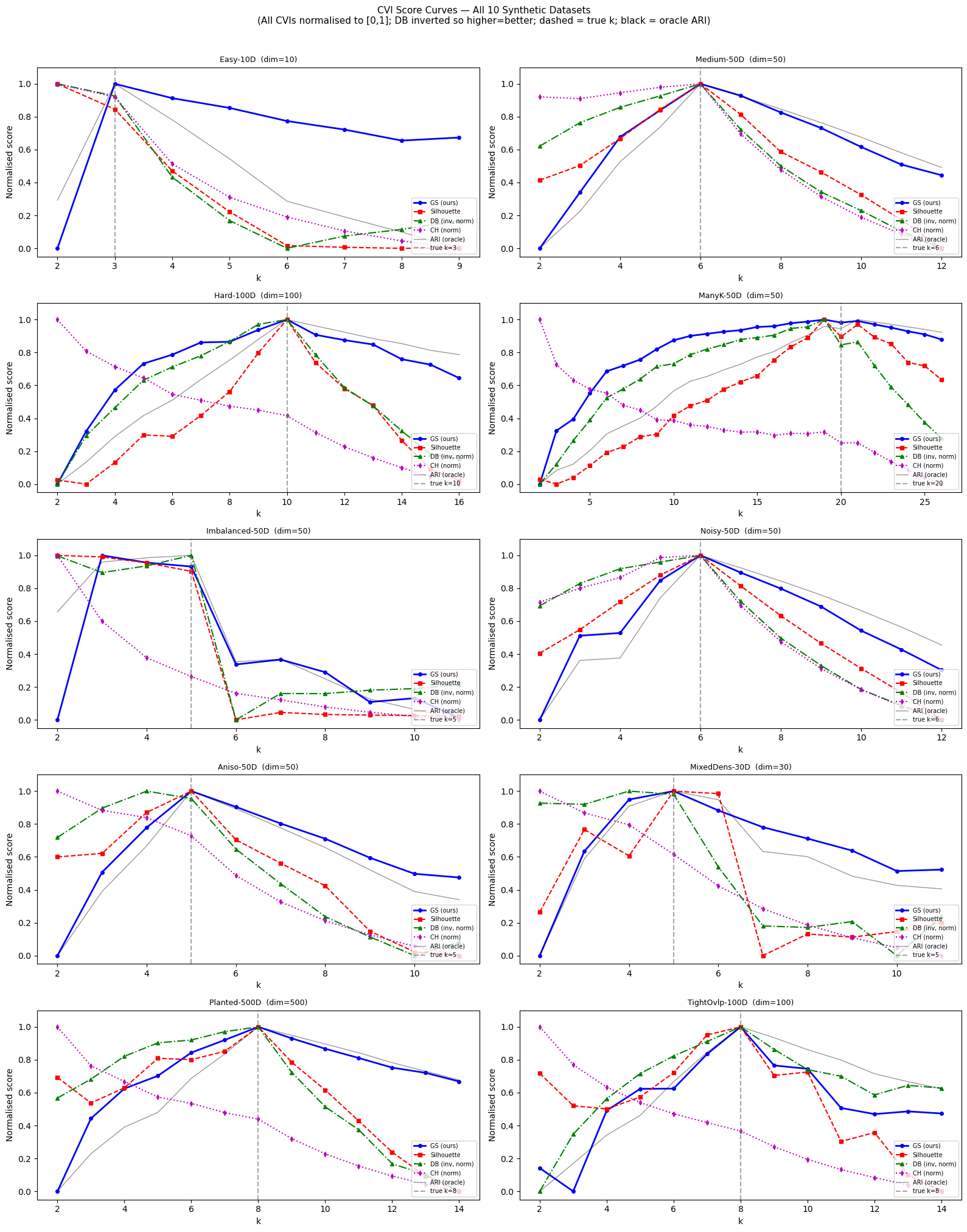
Where Silhouette collapses, Graph-SCOPE holds.
Silhouette is the standard unsupervised clustering metric used in scikit-learn, BERTopic, and thousands of production ML pipelines. It is also one of the most well-known examples of a metric that fails in high dimensions because it relies on pairwise Euclidean distances. The plot below shows Kendall's τ — the rank correlation between each metric's score and the true clustering quality — as data dimensionality grows from 2 to 5,000.