It should have been measuring structure.
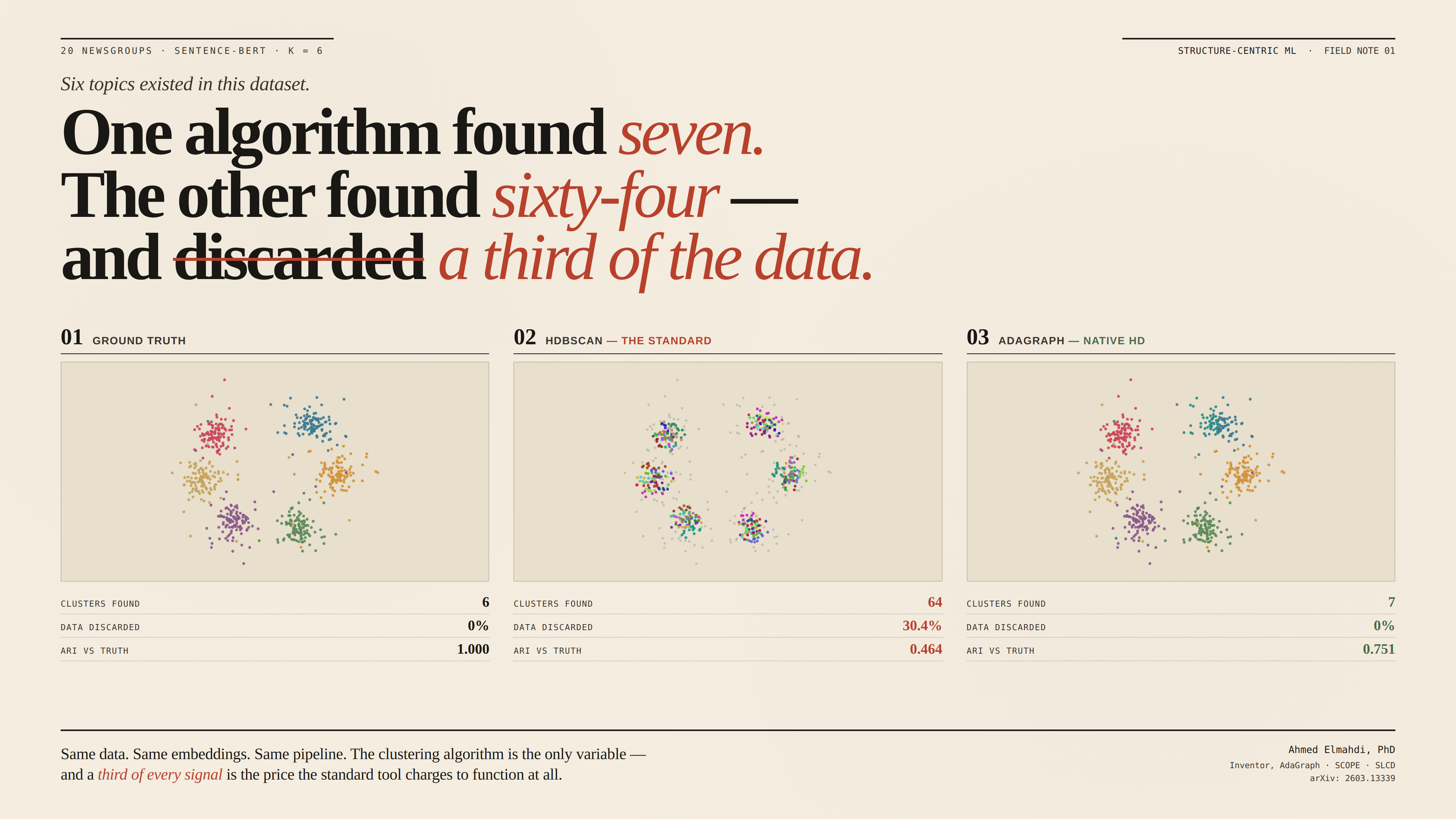
Structure-Centric Machine Learning is a new paradigm for unsupervised data analysis. It resolves three problems that DBSCAN (1996) and HDBSCAN (2013) never could: clustering in native high dimensions without reduction, retaining every data point instead of discarding noise, and transferring tuned parameters across dataset scales without re-tuning.