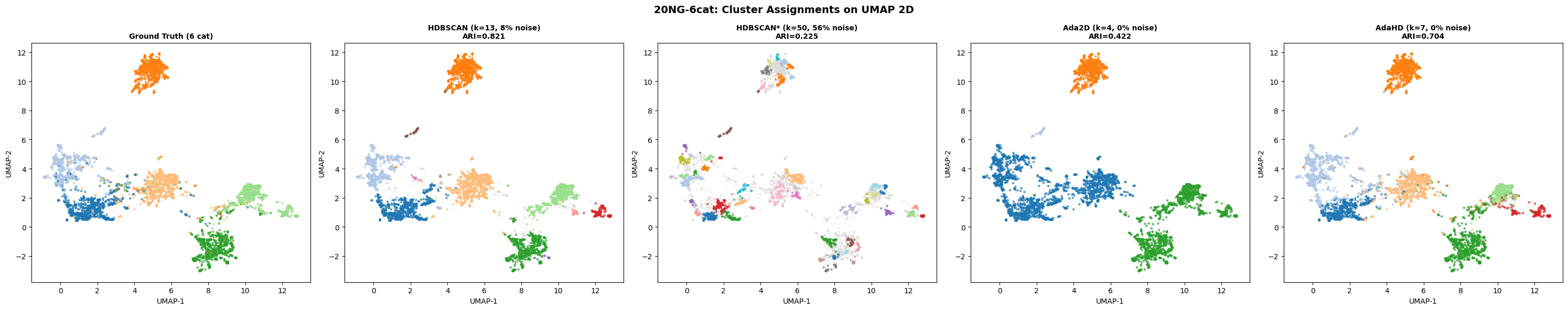
BERTopic pipeline, four datasets, four methods.
The standard text clustering pipeline in production today is BERTopic: sentence-BERT embeddings → UMAP to two dimensions → HDBSCAN. This is the pipeline that powers most social listening platforms, trend detection systems, and topic modeling tools. The benchmark below replaces only the final clustering step. Ada2D and AdaHD are scored on the same UMAP-reduced 2D data that HDBSCAN operates on.
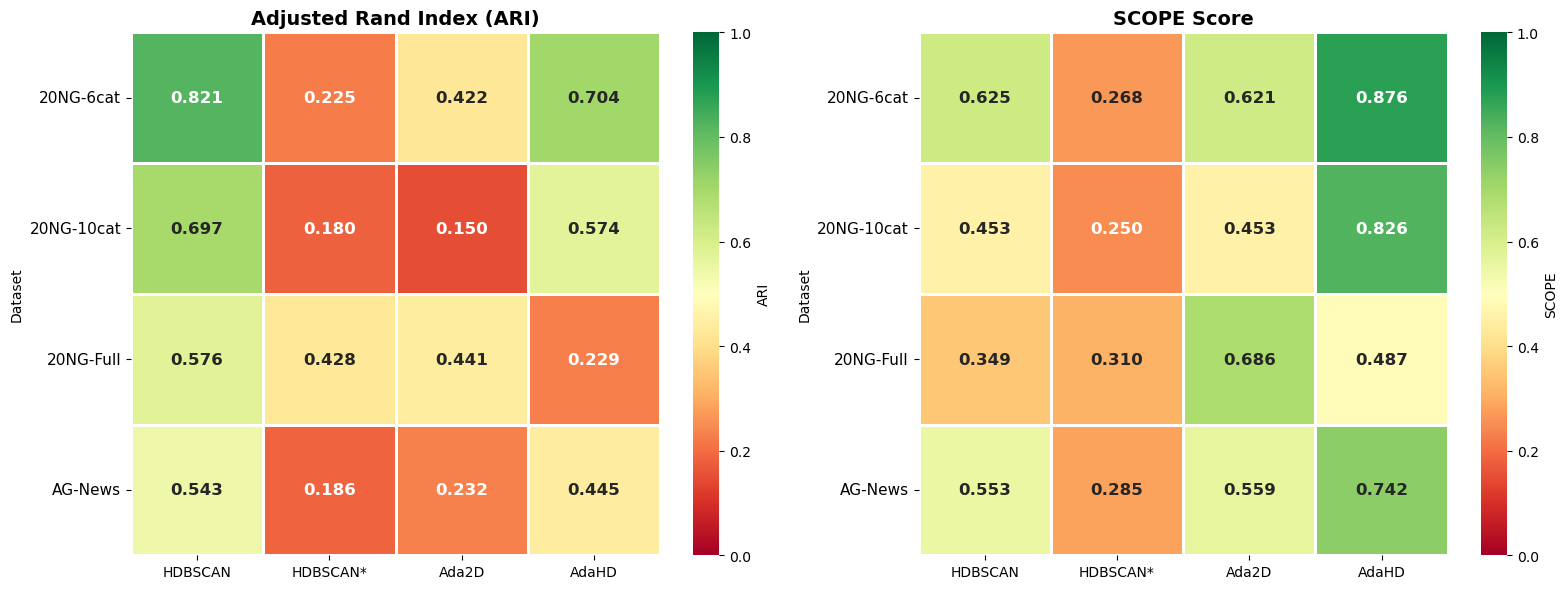
Average across all four datasets
| Method | Avg ARI | Avg SCOPE | Style |
|---|---|---|---|
| AdaHD (high-D structure-centric) | 0.5015 | 0.7604 | Native HD |
| Ada2D (low-D structure-centric) | 0.4261 | 0.5766 | UMAP → AdaBox |
| HDBSCAN* (tuned) | 0.3611 | 0.3595 | UMAP → HDBSCAN, hyper-tuned |
| HDBSCAN (default) | 0.3516 | 0.3017 | UMAP → HDBSCAN, default |